DICOM Viewer MCP
Interface UIpar thalesmms
Visualisez des séries d'images médicales DICOM directement dans Claude Desktop — navigation entre coupes, panoramique/zoom et affichage des métadonnées sans quitter votre interface IA.
Captures d'écran
Ce qu'il fait
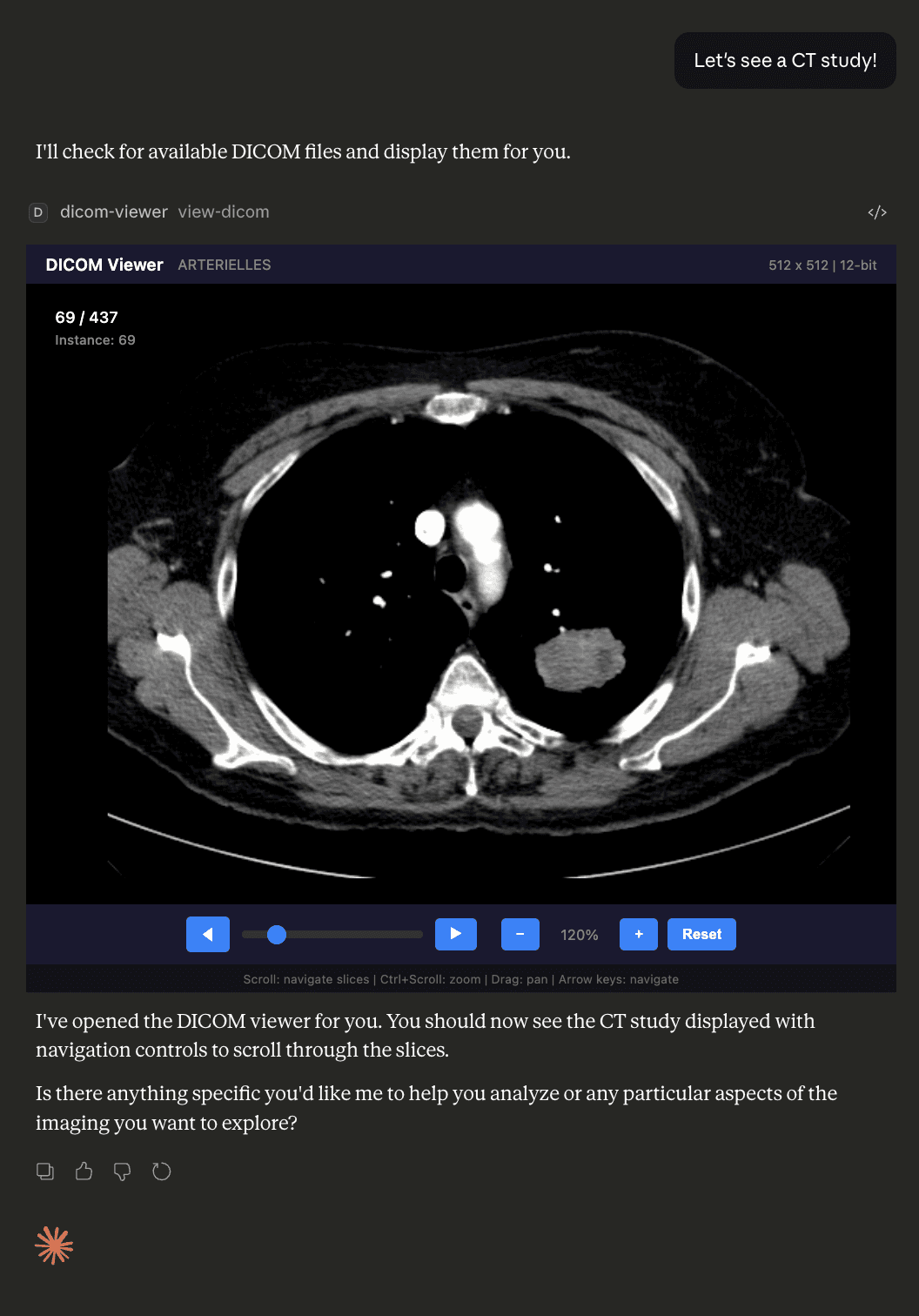
DICOM Viewer MCP est une application prototype MCP qui vous permet de charger et de naviguer dans des études d'imagerie médicale (fichiers .dcm) directement dans Claude Desktop. Il traite les fichiers DICOM côté serveur via dicom-parser et sharp, convertit les données de pixels en PNG avec une gestion appropriée du fenêtrage/niveau, et sert le résultat sous forme de ressource HTML interactive — contournant entièrement les restrictions de la Politique de Sécurité du Contenu (CSP).
Fonctionnalités clés
- Rendu DICOM intégré — aucun visionneur externe requis ; les images apparaissent dans Claude Desktop
- Navigation dans les séries — molette de défilement, curseur, boutons précédent/suivant et raccourcis clavier Début/Fin pour se déplacer entre les coupes
- Panoramique et zoom — Ctrl+molette pour zoomer, cliquer-glisser pour panoramer, bouton de réinitialisation pour restaurer la vue par défaut
- Traitement côté serveur — gère le fenêtrage/niveau, la pente/l'intercept de mise à l'échelle, MONOCHROME1/2, et les données de pixels 8-bit et 16-bit
- Tri automatique des coupes — trié par numéro d'instance ou position de la coupe
Installation
Clonez le dépôt et compilez localement, puis enregistrez-le dans votre configuration Claude Desktop.
git clone https://github.com/ThalesMMS/dicom-viewer-mcp-prototype.git
cd dicom-viewer-mcp-prototype
npm install
npm run build
Claude Desktop : Ajoutez à ~/Library/Application Support/Claude/claude_desktop_config.json (macOS) :
{
"mcpServers": {
"dicom-viewer": {
"command": "node",
"args": ["/absolute/path/to/dicom-viewer-mcp-prototype/dist/index.js", "--stdio"]
}
}
}
Placez les fichiers .dcm dans le dossier ./dicom/, redémarrez Claude Desktop, puis demandez à Claude de "montrer une étude DICOM".
Hôtes supportés
- Claude Desktop (transport stdio)
Note : Actuellement, seuls les fichiers DICOM non compressés sont supportés (Explicit/Implicit VR Little Endian). Les fichiers compressés JPEG/JPEG2000 ne sont pas encore supportés.